Project Documentation
Welcome to the documentation for this project. This page provides more background information about the technology and usage of this website.
Table of Contents
Introduction
This application was designed to gather and share knowledge about RNA polymerases in plants. It provides curated datasets to help researchers in the field prepare experiments, explore sequences or study phylogeny.
The system is built using Flask and SQLAlchemy for the backend and database management, with JavaScript for dynamic visualizations.
Getting Started
What can be found on polymerase.eu?
What cannot be found on polymerase.eu?
Database Organization
How is the data organized?
Unlike other databases, this one is structured around the protein sequence and everything is constructed around it. Each protein sequence is associated to a given isolate (variant, ecotype...) corresponding to a species. The combination of sequence and isolate is the main unit upon which the data is organized. The sequence itself (amino acid string) is linked to a unique identifier, a symbol, and a variant designation if applicable. For this protein, you'll see which RNA polymerase complexes it belongs to (Pol I through Pol VI) and its subunit number within those complexes. Additionally, you'll find database accessions (like Phytozome, UniProt, etc.) associated with this sequence across different biological databases, as well as any alternative names or symbols used in the scientific literature. This organization allows you to easily access all relevant biological context and cross-references for any given protein sequence, as well as visualize the state of the knowledge for a given species.
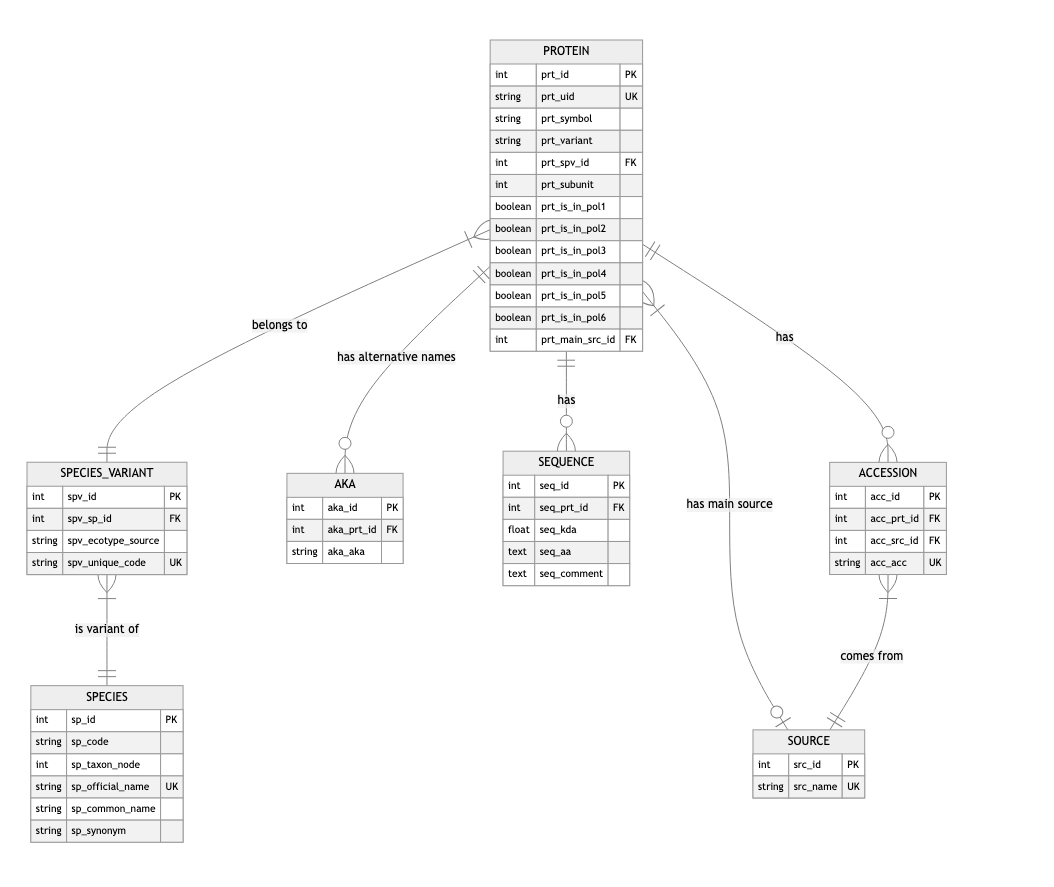
Table Relationships
The following diagram illustrates the relationships between tables.
Diagram